Which Machine Learning Classifiers are Best for Small Datasets…
An Empirical Study
Although "big data" and "deep learning" are dominant, my own work at the Gates Foundation involves a lot of small (but expensive) datasets, where the number of rows (subjects, samples) is between 100 and 1000. For example, detailed measurements throughout a pregnancy and subsequent neonatal outcomes from pregnant women. A lot of my collaborative investigations involve fitting machine learning models to small datasets like these, and it's not clear what best practices are in this case.
Along with my own experience, there is some informal wisdom floating around the ML community. Folk wisdom makes me wary and I wanted to do something more systematic. I took the following approach:
- Get a lot of small classification benchmark datasets. I used a subset of this prepackaged repo. The final total was 108 datasets. (To do: also run regression benchmarks using this nice dataset library.)
- Select some reasonably representative ML classifiers: linear SVM, Logistic Regression, Random Forest, LightGBM (ensemble of gradient boosted decision trees), AugoGluon (fancy automl mega-ensemble).
- Set up sensible hyperparameter spaces.
- Run every classifier on every dataset via nested cross-validation.
- Plot results.
All the code and results are here: https://github.com/sergeyf/SmallDataBenchmarks
Feel free to add your own algorithms.
Let's look at the results. The metric of interest is weighted one-vs-all area under the ROC curve, averaged over the outer folds.
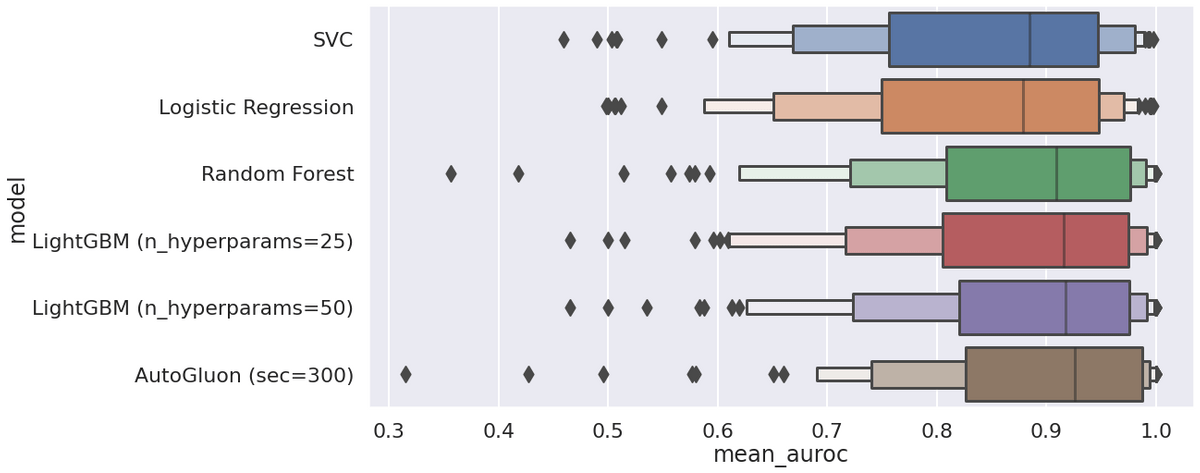
Some observations:
- AutoGluon is best overall, but it has some catastrophic failures (AUROC < 0.5) that Logistic Regression does not and LightGBM has fewer of.
- You can't tell from this particular plot, but AutoGluon needs "enough" time. It has a budget parameter which tells it how much time to spend improving the fancy ensemble. Five minutes per fold was the minimum that worked well - this adds up to 108 datasets * 4 outer folds * 300s = 1.5 days for the entire benchmark.
- Linear SVC is better than Logistic Regression on average. There are also two datasets where SVC is 0.3 and 0.1 AUROC better than every other model. It's worth keeping in the toolbox.
- Logistic Regression needs the "elasticnet" regularizer to ensure it doesn't have the kind of awful generalization failures that you see with AutoGluon and Random Forest.
- LightGBM is second best. I used hyperopt to find good hyperparameters. I also tried scikit-optimize and Optuna, but they didn't work as well. User error is possible.
- Random Forest is pretty good, and much easier/faster to optimize than LightGBM and AutoGluon. I only cross-validated a single parameter for it (depth).
Here are counts of datasets where each algorithm wins or is within 0.5% of winning AUROC (out of 108):
- AutoGluon (sec=300): 71
- LightGBM (n_hyperparams=50): 43
- LightGBM (n_hyperparams=25): 41
- Random Forest: 32
- Logistic Regression: 28
- SVC: 23
And average AUROC across all datasets:
- AutoGluon (sec=300): 0.885
- LightGBM (n_hyperparams=50): 0.876
- LightGBM (n_hyperparams=25): 0.873
- Random Forest: 0.870
- SVC: 0.841
- Logistic Regression: 0.835
And counts where each algorithm does the worst or is within 0.5% of the worst AUROC:
- Logistic Regression: 54
- SVC: 48
- Random Forest: 25
- LightGBM (n_hyperparams=25): 19
- LightGBM (n_hyperparams=50): 18
- AutoGluon (sec=300): 14
Which shows that even the smart ensemble can still fail 10% of the time. Not a single free lunch to be eaten anywhere.
Here is a plot of average (over folds) AUROC vs number of samples:
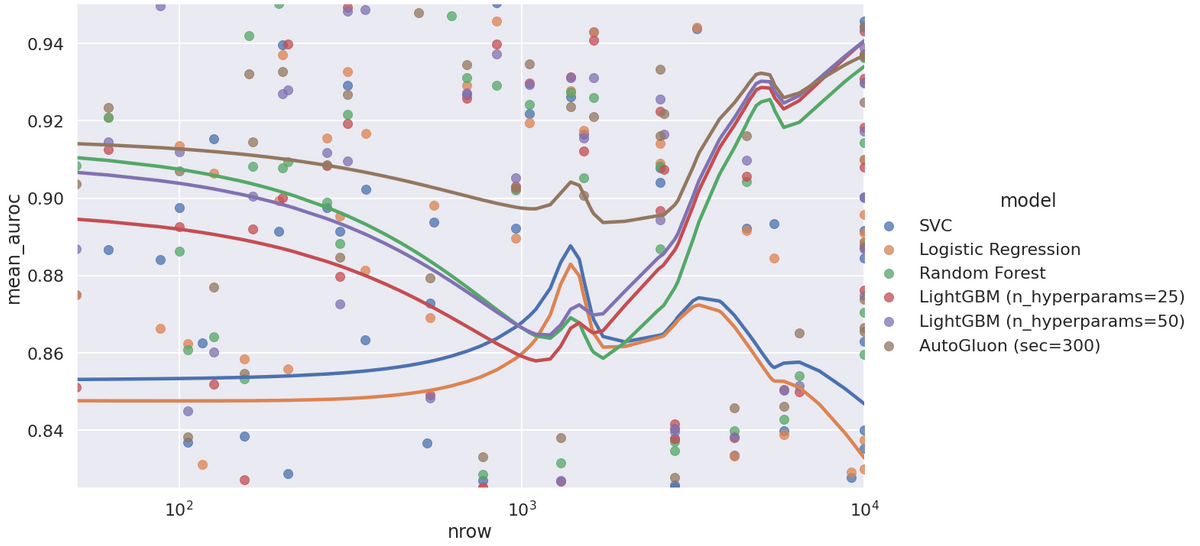
I was surprised when I saw this plot the first time. The collective wisdom that I've ingested is something like: "don't bother using complex models for tiny data." But this doesn't seem true for these 108 datasets. Even at the low end, AutoGluon works very well, and LightGBM/Random Forest handily beat out the two linear models. There's an odd peak in the model where the linear models suddenly do better - I don't think it's meaningful.
The last plot - standard deviation of AUROC across outer folds:
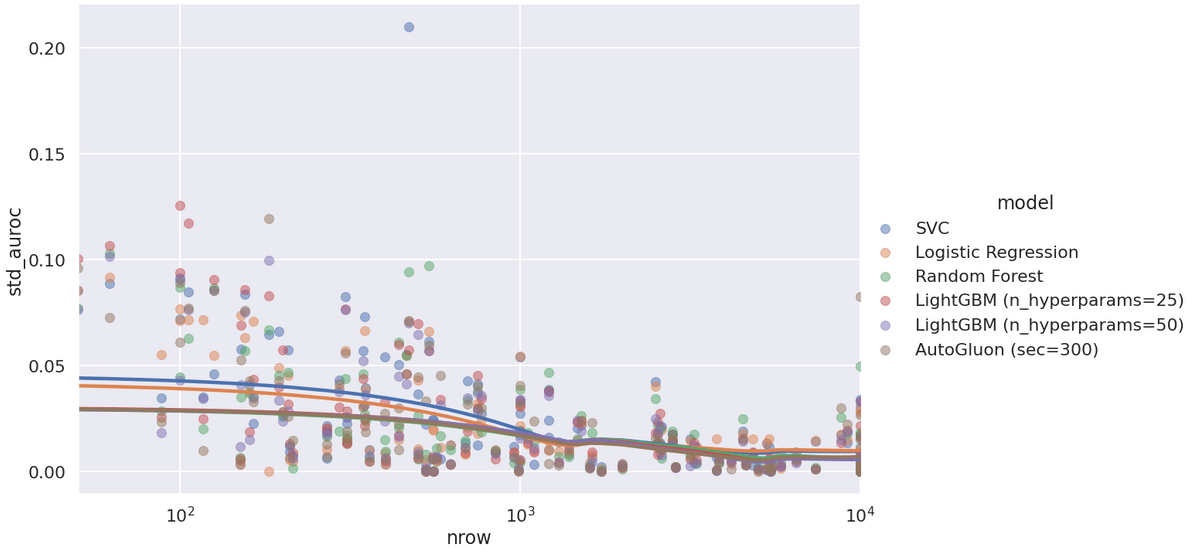
Linear models don't just generalize worse regardless of dataset size - they also have higher generalization variance. Note the one strange SVC outlier. Another SVC mystery...
IID Thoughts
How applicable are these experiments… Both levels of the nested cross-validation used class-stratified random splits. So the splits were IID: independent and identically distributed. The test data looked like the validation data which looked like the training data. This is both unrealistic and precisely how most peer-reviewed publications evaluate when they try out machine learning. (At least the good ones.) In some cases, there is actual covariate-shifted "test" data available. It's possible that LightGBM is better than linear models for IID data regardless of its size, but this is no longer true if the test set is from some related but different distribution than the training set. I can't experiment very easily in this scenario: "standard" benchmark datasets are readily available, but realistic pairs of training and covariate-shifted test sets are not.
Conclusions & Caveats
So what can we conclude…
- If you only care about the IID setting or only have access to a single dataset, non-linear models are likely to be superior even if you only have 50 samples.
- AutoGluon is a great way to get an upper bound on performance, but it's much harder to understand the final complex ensemble than, say, LightGBM where you can plot the SHAP values.
- hyperopt is old and has some warts but works better than the alternatives that I've tried. I'm going to stick with it.
- SVC can in rare cases completely dominate all other algorithms.
Caveats:
- LightGBM has a lot of excellent bells and whistles that were not at all used here: native missing value handling (we had none), smarter encoding of categorical variables (I used one-hot encoding for the sake of uniformity/fairness), per-feature monotonic constraints (need to have prior knowledge).
- AutoGluon includes a tabular neural network in its ensemble, but I haven't run benchmarks on it in isolation. It would be interesting to find out if modern tabular neural network architectures can work out-of-the-box for small datasets.
- This is just classification. Regression might have different outcomes.
Again, check out the code and feel free to add new scripts with other algorithms. It shouldn't be too hard. https://github.com/sergeyf/SmallDataBenchmarks
